Use of RNA profiles to predict therapeutic response to standard treatment and for the identification of potentially novel therapeutic options in colorectal adenocarcinoma
Colorectal carcinoma (CRC) is one of the most frequent cancers. Approximately every sixteenth woman and every nineteenth man are diagnosed with CRC during their lifetime in Europe.
The standard treatments after surgery and besides radiation-therapies are chemotherapies and targeted therapies. For targeted therapies, distinct biomarkers such as overexpressed proteins or specific mutations are used for the decision of the therapy. However, many patients are negative for these biomarkers and receive chemotherapy, often in different combinations. Which patient responds well or not to which chemotherapy or combination of drugs is unknown and new biomarkers are urgently needed to make better patient-specific therapeutic decisions.
For this purpose, GenXPro is the partner in REVERT with the particular role of examining if the therapeutic decision can be supported by information obtained from RNA-data from CRC patients. RNA from cancer tissue is usually heavily degraded and can often not be measured accurately. In order to measure the gene expression of the biopsies that are generated from Formalin-Fixed Paraffin-Emebedded (FFPE) biopsies of the tumors, GenXPro used “MACE-Seq” a 3’mRNA method that is highly sensitive and does not require high-quality RNA. In addition to the mRNA analyses from poly-adnalyted transscripts which are usually protein coding, Genxpro examined as well smallRNAs from the patient derived FFPE samples using our proprietary highly sensitive “TrueQuant smallRNA Kit”.
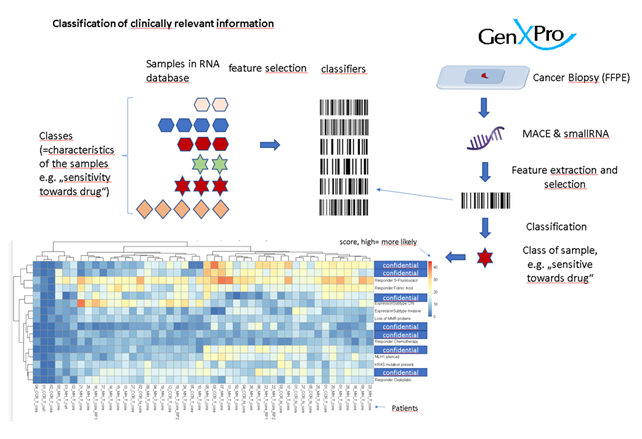
The RNA-profiles that were obtained were compared to RNA profiles from CRC patients that are available in public databases and which contained therapeutically relevant information such as survival and response to the different treatments. In order to identify the characteristic gene expression features that are responsible for a certain trait such as sensitivity to a drug, a classifier-database was generated using machine-learning and statistical approaches.
In addition, RNA-data from cell cultures which contained information about the sensitivity of the cell cultures towards numerous drugs was used to identify potentially alternative drugs for the treatment of CRC. Moreover, well established information such as the consensus molecular subtype (CMS) which is used for the grading of the CRC were generated using MACE-Seq data.
The information about the sensitivity towards other drugs was collected in a report for each patient (“ClinXPro-report”). In a collaboration with Pablo Conesas group in Valencia (Spain), patient derived organoids (PDOs) were generated. PDOs are an emerging tool which allow the identification of mechanisms underlying CRC pathogenesis. In this study, we examined CRC-PODs representing MC and SAC to characterize these rare subtypes and determine their CMS, as well as to perform the in silico drug sensitivity assay (ClinXPro report based on MACE-Seq).
A special focus was also set on the better understanding between medullary cancer (MC), serrated (serrated adenocarcinoma, SAC). and non-serrated CRC.
Moreover, GenXPro examined healthy tissue, primary tumors and metastatic tumors of the same patient in order to obtain insights into the development mechanisms from primary to metastatic CRCs.
All analyses were performed using the above described ClinXPro analyses in addition to pathway- enrichment and GO enrichment analyses.