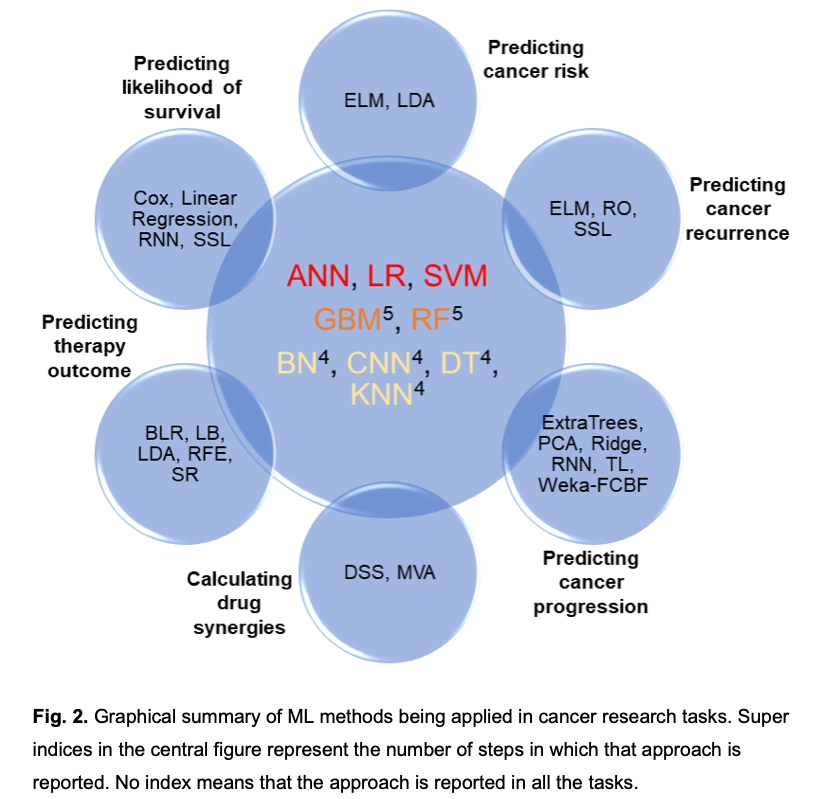
Artificial Intelligence is providing astonishing results, with medicine being one of its favourite playgrounds. In the short-term future, computers may be capable of formulating diagnoses and choosing the correct treatment, while robots may perform surgical operations, and conversational agents could interact with patients as virtual coaches. Machine Learning and, in particular, Deep Neural Networks are behind this revolution. In this scenario, important decisions will be controlled by standalone machines that have learned predictive models from provided data. Among the most challenging targets of interest in medicine are cancer diagnosis and therapies but, to start this revolution, software tools need to be adapted to cover the new requirements. In this sense, learning tools are becoming a commodity in programming libraries such as Python and Matlab, just to name two, but to exploit all their possibilities, it is essential to fully understand how models are interpreted and which models are more interpretable than others. We focus on the analysis of current machine learning models, frameworks, databases and other related software tools as applied to medicine – specifically, to cancer research – and we discuss their interpretability, computational performance and the input data they ingest. Amongst the different types of cancer, we have focused on colorectal cancer (CRC) because it is the fourth cause of mortality worldwide and it is a remarkable worry for society. From the evidence available, artificial neural networks (ANN), logistic regression (LR) and support vector machines (SVM) have been observed to be the preferred models. Besides, convolutional neural networks (CNN) which are a subtype of ANN typically used for image processing, supported by the rapid development of GPUs and tensor-oriented programming libraries, are gaining in importance. However, the interpretability of results by doctors is rarely considered which is a factor that needs to be improved. We therefore consider this study to be a timely contribution to the issue.
Click here for the arxiv pre-print:https://arxiv.org/abs/2010.00353